DATE2023.04.06 #Press Releases
Study reveals the 3D structure of a protein involved in genome editing
Understanding the evolution of the CRISPR-Cas enzyme from the TnpB protein will provide new insights into targeted genome editing technology.
April 06, 2023
Gene editing is one of the latest breakthroughs in biology. The widely known CRISPR-Cas gene editing system provided prokaryotes (organisms that lack cell nuclei) an immunity against foreign DNA. Since the discovery of the CRISPR gene editing technology, scientists are revealing the evolution of the CRISPR-Cas proteins from their precursors. That knowledge will help them develop other small and new genome editing tools for gene therapy. At the University of Tokyo, Prof. Osamu Nureki’s group works on identifying the structure and function of the proteins involved in genome editing. In a recent study by the team, they discovered the 3D structure of a protein called TnpB, a probable precursor to the CRISPR-Cas12 enzyme. Their findings appeared in the journal Nature.
Previous research suggests that the TnpB protein may work like a pair of molecular scissors, cutting DNA with the help of a special non-coding RNA called ωRNA. But exactly how the RNA-guided DNA cleavage works and its evolutionary relationship with Cas12 enzymes was unknown, which prompted Nureki lab’s investigation. Their first and most crucial step toward understanding that was to reveal the protein structure.
To determine the three-dimensional structure of TnpB, the researchers took the protein TnpB from a bacterium called Deinococcus radiodurans and used cryo-electron microscopy. In the cryo-electron microscopy technique, the protein sample is cooled to -196° C using liquid nitrogen and irradiated with electron beams, which reveals the protein’s 3D structure (Figure 1). The team found that the ωRNA in TnpB has a unique pseudoknot shape similar to that found in the guide RNAs of Cas12 enzymes. The study also revealed how TnpB recognizes the ωRNA and cuts the target DNA. And when they compared the structure of this protein to Cas12 enzymes, they learned two possible ways TnpB might have evolved into CRISPR-Cas12 enzymes (Figure 2).
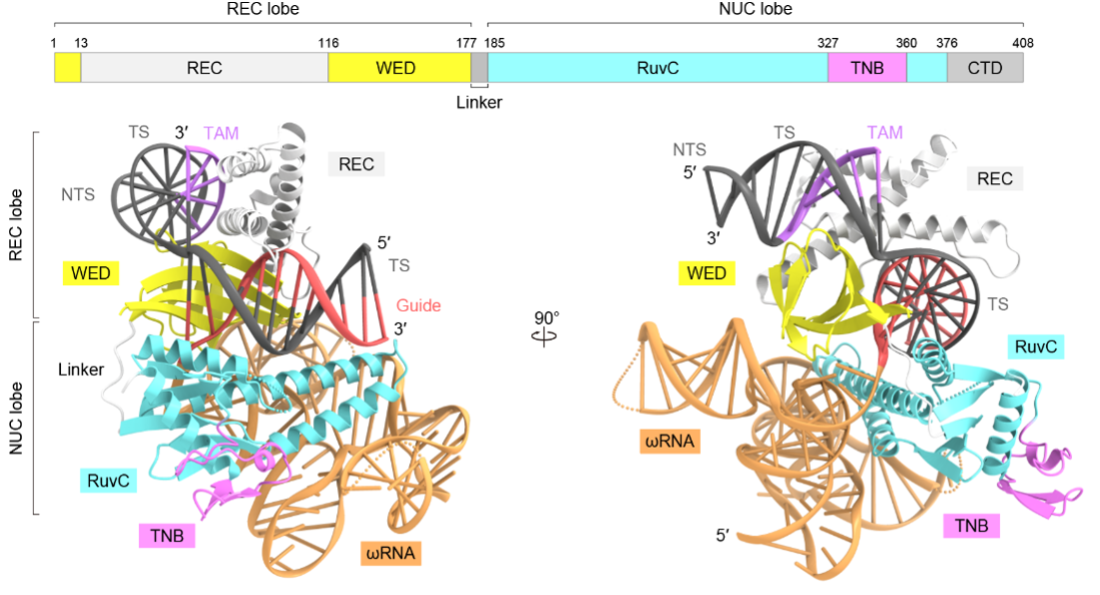
Figure 1: Overall structure of TnpB-ωRNA-target DNA complex. TnpB adopts a bilobed structure consisting of the REC lobe and NUC lobe. The REC lobe comprises WED and REC domains, whereas the NUC lobe comprises the RuvC and TNB domains. TnpB assembles with a ωRNA transcribed from the transposon sequence at the end of its coding region. The guide RNA-target DNA heteroduplex is bound to the groove between the REC and NUC lobes.
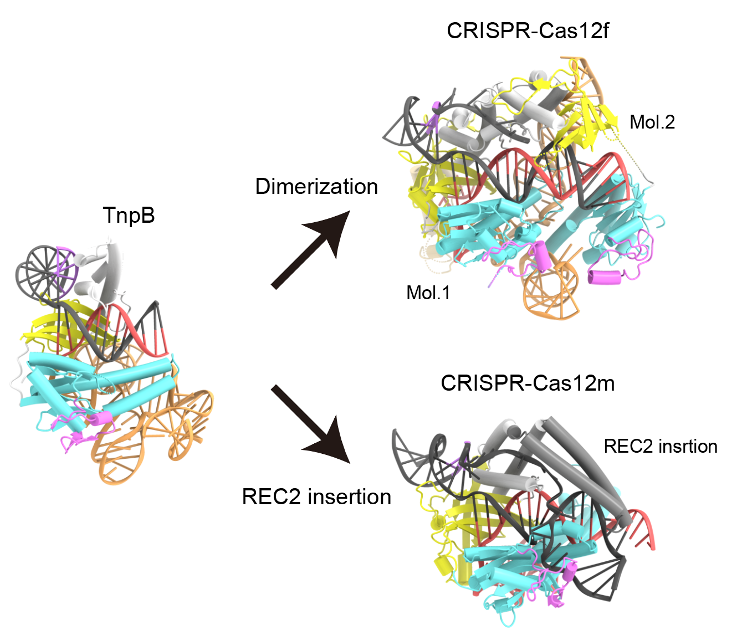
Figure 2: Evolutionary path from TnpB to Cas12 enzymes. Structural comparison of TnpB with Cas12 enzymes with high sequence similarity to TnpB revealed that Cas12 enzymes recognize longer target DNA sequences than TnpB by either asymmetric dimer formation or diverse REC2 insertions, allowing for engagement in CRISPR-Cas adaptive immunity.
“Our findings provide mechanistic insights into the TnpB function and advance our understanding of the evolution from TnpB proteins to CRISPR-Cas12 effectors,” says Ryoya Nakagawa, a graduate student and one of the first authors of the research paper. In the future, he added, “We will explore the potential applications of TnpB-based gene editing techniques.”
Publication details
Journal Nature Title Cryo-EM structure of the transposon-associated TnpB enzyme AuthorsRyoya Nakagawa, Hisato Hirano, Satoshi N. Omura, Suchita Nety, Soumya Kannan, Han Altae-Tran, Xiao Yao, Yuriko Sakaguchi, Takayuki Ohira, Wen Y. Wu, Hiroshi Nakayama, Yutaro Shuto, Tatsuki Tanaka, Fumiya K. Sano,
Tsukasa Kusakizako, Yoshiaki Kise, Yuzuru Itoh, Naoshi Dohmae, John van der Oost, Tsutomu Suzuki, Feng Zhang & Osamu NurekiDOI


