DATE2023.01.12 #Press Releases
Machine learning predicts bacterial genome evolution
Bacterial metabolic system evolution is predictable and has implications for antibiotic resistance and bioengineering.
January 12, 2023
Naoki Konno and Wataru Iwasaki from the University of Tokyo have developed a bioinformatics method called Evodictor which can predict future bacterial evolution.
Can we predict evolution? Some argue that we cannot because random effects drive it; others reason that evolutionary pressures and constraints make it predictable. Previous research has provided some evidence for the latter. But the evidence is limited to short-term, DNA sequence-level evolution. In a study published in the journal Science Advances, Konno and Iwasaki demonstrate that the long-term evolution (gene gain or loss) of bacterial metabolic systems is predictable.
“The predictability of evolution itself is a critical problem in biology because evolution is the principal power that enables the existence of all life systems on earth. In addition, the prediction of gene gains or losses by Evodictor is expected to contribute to solving problems in society. In medical fields, there is an increasing demand for ranking species according to the risk of gaining antimicrobial resistance genes by dissemination from other species. In bioengineering and industrial fields, there is a strong requirement for computationally screening species to which we can readily add or delete specific genes before conducting costly experiments,” says Prof. Iwasaki.

Figure 1. The framework of Evodictor. Evodictor predicts the probability of gene gain or loss from a gene repertoire of genomes (i.e., presence or absence of every gene) by learning past gene gain or loss evolution across diverse species.
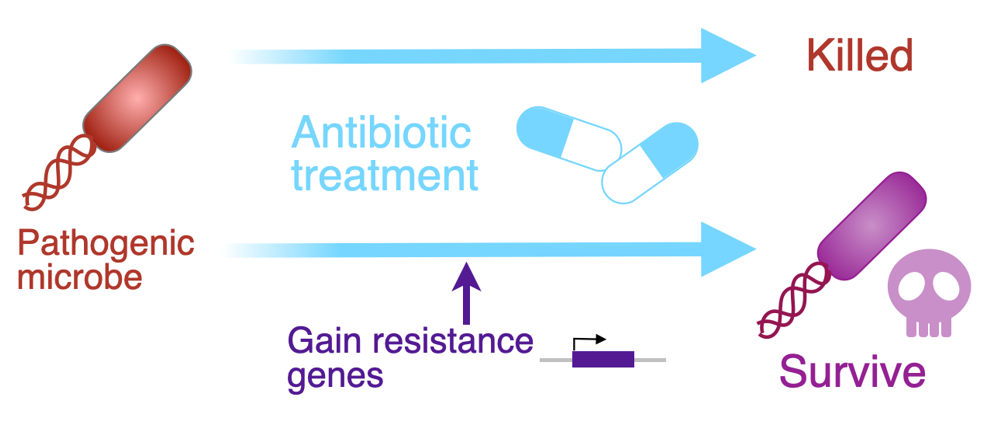
Figure 2. Antibiotic resistance evolution. Microbes can survive against antibiotics after acquiring resistance genes, which is a worldwide problem in the medical field now.
The duo used large-scale data from around 3000 bacterial genomes and machine learning to predict the evolution of bacterial metabolic genes (Figure 1). “First, we adopted a mathematical model of gene gain or loss evolution and reconstructed gene repertoires of the past ancestral species on a phylogenetic tree of life. This lets us collect large-scale data on the long-term evolution of biological systems across diverse species, which are hardly observable in laboratories. Second, we computationally extracted characteristic patterns within that reconstructed evolutionary history by machine learning, a powerful computational framework that recently attracted great publicity as artificial intelligence. Finally, the machine learning model employed the extracted patterns to predict the gain or loss probability of every gene,” explains Konno.
The machine learning model could predict metabolic gene evolution (both gene gains and losses) across various bacterial species. The finding implies that the evolutionary pressures and constraints on the metabolic systems are similar across bacteria. So similar evolutionary patterns will be observed in distinct species, which can be used to predict future evolution. In the future, the duo hopes to experimentally verify the evolutionary patterns predicted by machine learning. They also envision predicting the probability of gaining antimicrobial resistance genes across various pathogens. Such a database would prove immensely useful for our fight against antibiotic resistance (Figure 2).
“Finding rules and patterns behind the life system evolution is the critical path to answering why we are here, what future organisms will look like, and how we can fully harness the power of bioengineering technologies,” says Prof. Iwasaki. For instance, predicting the evolutionary patterns of pathogens will provide insights into designing drugs that avoid antibiotic resistance evolution. Predicting cancer cell evolution can help diagnose cancer stages and develop effective treatments.
Publication details
Journal Science Advances Title Machine learning enables prediction of metabolic system evolution in bacteriaAuthors DOI


